Blog Posts
Publication: collaborative paper on perovskite-related materials, with Sn and Se ssNMR
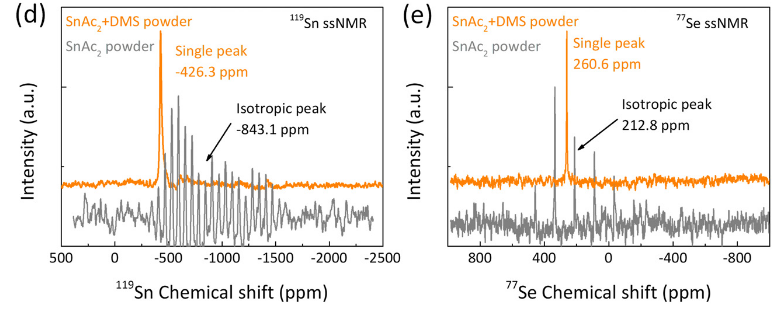
Congrats to our collaborators from the group of Maria Loi and others for the publication of their new paper on in situ SnSe deposition as passivation for scalable and stable quasi-2D lead–tin perovskite solar cells, in the journal Energy & Environmental Science. Dr. Lasorsa in our group contributed tin and selenium ssNMR analysis to this interdisciplinary paper. For more information see the paper at the journal.
Publication:
Chen L, Tekelenburg EK, Gahlot K, Pitaro M, Xi J, Lasorsa A, et al. In situ SnSe deposition as passivation for scalable and stable quasi-2D lead–tin perovskite solar cells. Energy Environ Sci. 2023;10.1039.D3EE02507A.
Upcoming Symposium Dutch Protein Aggregation Network DPAN
Hold the date! On November 30th there will be an inaugural symposium of the Dutch Protein Aggregation Network (DPAN), sponsored by NWO. This one-day even (free registration) will be held at the VU in Amsterdam. It welcomes all Dutch researchers interested in protein aggregation related to human disease (including PIs, PhD students, postdocs..).
The DPAN network aims to bring together the protein aggregation researchers of The Netherlands. With the help of NWO sponsorship DPAN will start with a one-day symposium on Nov 30th. This will take place at the O|2 building of the VU in Amsterdam.
The keynote speaker will be Harm Kampinga from the UMCG Groningen. Many other speakers will discuss their research on amyloid formation and other aspects of protein aggregation.
Please join us and register here.
Educational webinar on the use of ssNMR to measure dihedral angles.
Recently, Patrick gave an online lecture in the online webinar tutorial series of the Global NMR Discussion Meetings series (episode 70!). In this online lecture he introduced and discussed approaches to measure torsion angles (or dihedral angles) using advanced solid-state NMR spectroscopy. He discussed the basic idea of how these are implemented in ssNMR, but also how such structural data can be a useful complement to more traditional inter-atomic distance information obtained by ssNMR. You can now watch the recording of this lecture on Youtube, via this link.
Want to learn even more?
This lecture is connected to our recent review article on the same topic, which can be found online at: Frontiers | Dihedral Angle Measurements for Structure Determination by Biomolecular Solid-State NMR Spectroscopy (frontiersin.org)
Publication: 13C labeling of hyaluronic acid polysaccharides (for ssNMR analysis)
Congrats to Pushpa and her collaborators on the new paper in the journal Carbohydrate Polymers. This new report describes Pushpa’s work to achieve the production of high-molecular-weight (HMW) hyaluronic acid, as part of her PhyCan (physics of cancer) project. Our interest in this polysaccharide stems from its important role in the extracellular matrix (ECM) of tissues, and in particular certain types of cancer tissues. In such tumors the amount of HA is upregulated, seemingly contributing to the progression of cancer development. A challenge in understanding this process is that HMW is difficult to study with most structural techniques, as it can be very large (megadaltons!) and highly dynamic. In this project, Pushpa outfitted HMW HA with 13C (and 15N) isotope labels, and shows that this makes multidimensional NMR (including solid-state NMR) feasible. This approach is expected to be valuable for many research areas, as HA is also important for many biomedical engineering (BME) type applications. ECM-mimicking hydrogels are for instance of great interest for engineering 3D environments for growing cells.
Publication:
Rampratap P, Lasorsa A, Perrone B, Van Der Wel PCA, Walvoort MTC. Production of isotopically enriched high molecular weight hyaluronic acid and characterization by solid-state NMR. Carbohydrate Polymers. 2023 Sep;316:121063.
Job search: NMR spectroscopist being recruited at the RuG
The University of Groningen is looking to hire a NMR spectroscopist for the solution NMR facility associated with the chemistry-focused Stratingh Institute. Ad is here: https://www.rug.nl/about-ug/work-with-us/job-opportunities/?details=00347-02S0009SXP&cat=obp
Note that this position is not associated with our research group. The mentioned NMR facility is part of the Stratingh Institute and focuses on solution NMR studies.
Publication: Dynamics-based spectral editing to see (fiber) surfaces by solid-state NMR.
Congratulations to Dr. Irina Matlahov and (alum) Dr. Jennifer Boatz on the publication of their new paper in the Journal of Structural Biology X. The paper is entitled “Selective observation of semi-rigid non-core residues in dynamically complex mutant huntingtin protein fibrils“. It describes our latest research on the misfolded protein deposits associated with Huntington’s disease (HD), specifically looking at what happens on the surface of these protein fibrils.
In previous work we have studied the structure of these nanometer-sized fibrils formed by mutant huntingtin’s exon 1 fragment, using ssNMR, EM and other methods. In our earlier studies we used dynamics-sensitive NMR techniques to look at the core of these fibrils, taking advantage of its very rigid structure. Conversely, we had looked at super flexible protein parts that are away from the rigid core. Those types of techniques have been a popular tool for seeing rigid or flexible parts of (aggregated) proteins. However, in the current paper we try to look for protein parts with intermediate mobility. Of particular interest are those residues that form the surface of the rigid fibril core: residues that are somewhat immobilized by proximity to the core proper, but are somewhat mobilized by their interactions with water solvent surrounding the fibril. This part of the protein fibrils can be especially interesting, as it is the part seen by protein-targeting antibodies, PET ligands and protein-protein interaction partners in affected neurons.
The paper discusses how we can probe the impact of the water interactions with the surface through variable temperature ssNMR. However, it also shows the limitations of this traditional approach. Instead we advocate for a new type of dynamic filtering experiment that selectively shows the signals of semi-rigid/semi-mobile residues (such as those on the fiber surface). This technique (IMS-DYSE) is complementary to traditional DYSE methods that select rigid or flexible sites. For more details, please see the paper (linked below). We expect to combine this dynamic filtering technique with other types of pulse sequences, to probe in more detail the features of the fiber surface.
Note that this technique proved especially useful for our work on polyglutamine protein fibrils, as in this protein system we have a core made up of glutamine residues, providing an overwhelming strong NMR signal that masks the resonances of the surface glutamines. So, to see the surface, it is essential to find a way to suppress the core signals, leaving only those of the surface residues. The paper shows that this indeed works, and we can for the first time use this technique to distinguish the core and surface glutamines. With this technique in hand, we can foresee further studies of surface-specific features, such as the binding sites for targeted antibodies and/or amyloid-binding dyes.
The paper is available online at the journal, and has the following citation:
I. Matlahov, J.C. Boatz, P.C.A. van der Wel (2022) Selective observation of semi-rigid non-core residues in dynamically complex mutant huntingtin protein fibrils. J. Struct. Biol. X, vol. 6, 100077. DOI: 10.1016/j.yjsbx.2022.100077
PS. Some of the techniques and results from this publication were also discussed during a prior online seminar in the MIT ssNMR/DNP zoominar series. You can view that video here.
Publication: Collaborative paper with the Pescarmona group (ENTEG) on zeolites.
Congratulations to PhD student Mustapha El Hariri El Nokab and our collaborators from the Pescarmona group at ENTEG, on a new collaborative paper being accepted and posted online. In this work, Mustapha used both 29Si and 27Al magic angle spinning ssNMR to compare different zeolite samples, characterizing their chemical structure and degree of order. Aside from our ssNMR data, the paper features numerous other spectroscopic and synthetic methods. For more information, see the paper!
Zahra Asgar Pour, Romar Koelewijn, Mustapha El Hariri El Nokab, Patrick C. A. van der Wel, Khaled O. Sebakhy, Paolo Pescarmona (2022) Binder-free zeolite Beta beads with hierarchical porosity: synthesis and application as heterogeneous catalysts for anisole acylation. ChemCatChem in press.
News: Dutch-language radio
Recently our research was covered (briefly) on a Dutch radio station, featuring an interview with Patrick. It is mostly about our work applying ssNMR to studying how proteins aggregate in Huntington’s disease. You can find the recording online at the NPO radio website. More information about our Huntington disease research can also be found on this page, and in two recent webinars.

Upcoming event: BPS networking event on polyQ biophysics
Hereby an early announcement of an upcoming online event hosted by our group and the group of Markus Miettinen: “Biophysics of polyglutamine aggregation: how does it start and how does it end?“
This is an online event is organized in context of the networking event series sponsored and organized by the Biophysical Society (BPS). Our event aims to bring together international scientists working on the biophysics of polyglutamine (polyQ) protein aggregation. The event will take place on July 6th 2022, from 15:00-18:00 CET.
More information can be found at the page from the BPS or our own institute. An online registration page is planned to online at the BPS, see below. If you want more information in the mean time, please contact us by email.
Update May 2022: the registration page is now active at the BPS. The link is here. Note that this does require a free BPS account to work. Let us know if you have any issues or questions.
Publication: SSNMR of alginate hydrogel (re)hydration
A new open-access publication by Mustapha and collaborators has been published in the journal Food Hydrocolloids, describing how he used various ssNMR measurements to probe alginate hydrogel structure and (re)hydration. Alginates can be cross-linked with calcium to form hydrogels, which are used in many different types of applications. This includes their use in slow-release drug delivery as well as food/nutritional applications. In this new paper, Mustapha shows how 1H, 2H and 13C ssNMR measurements can be used to see how these hydrogels are structured, but especially also how they are hydrated by the aqueous solvent. A key feature of the work is the use of 1H MAS NMR to detect the water molecules in the hydrogel macropores as well as those waters directly interacting with the alginate. The two types of water molecules form distinct ‘pools’ in the hydrogel, which can be nicely distinguished by their different molecular motion. The latter is detected via different relaxation measurements, in these NMR experiments.
To read more:
El Hariri El Nokab, M..; Lasorsa, A.; Sebakhy, K. O.; Picchioni, F.; van der Wel, P. C. A. Solid-State NMR Spectroscopy Insights for Resolving Different Water Pools in Alginate Hydrogels. Food Hydrocolloids 2022, 107500.