news
Publication: New paper about photochemical approaches to studying polyQ protein aggregation.
Congratulations to PhD student Raffaella Parlato, Dr. Jana Volaric, and their collaborators, on the publication of a nice new paper in the Journal of the American Chemical Society. This is the final result of an idea from some years ago, which came together very nicely thanks to a great team of collaborators. The goal of this work was to explore the idea of putting polyglutamine aggregation under some degree of photo-control. In prior work, it has been shown that b-hairpin formation is a key step in the aggregation process of expanded polyQ proteins. Azobenzene-based groups can be used to favor or disfavor beta turn structures under the influence of light, but prior ways of implementing this have been limited in various ways. A new amino acid analogue is introduced with more favorable structural and photochemical properties. And, we tested it in the context of polyQ peptides, seeing it indeed modulating the conformation of resulting aggregates (seen by ssNMR on unlabeled peptide fibrils!). For more details, see the paper. It is open access, so easily accessible.
Citation:
Raffaella Parlato, Jana Volarić, Alessia Lasorsa, Mahdi Bagherpoor Helabad, Piermichele Kobauri, Greeshma Jain, Markus S. Miettinen, Ben L. Feringa*, Wiktor Szymanski*, and Patrick C. A. van der Wel* (2024) J. Am. Chem. Soc. 2024, 146, 3, 2062–2071. Photocontrol of the β-Hairpin Polypeptide Structure through an Optimized Azobenzene-Based Amino Acid Analogue. DOI: https://doi.org/10.1021/jacs.3c11155
Publication: Review article on polyglutamine protein analysis by solid-state NMR spectroscopy.
Our new review article summarizing progress in polyglutamine protein studies by solid-state NMR is now available on the website of the journal Biochemical Society Transactions. This is an invited mini-review that was requested to cover the progress made in our understanding the aggregation and misfolding of mutant polyglutamine proteins implicated in diseases such as Huntington’s disease and several versions of spinocerebellar ataxia (SCA). We describe how studies from ssNMR has been used to better understand the structure of these protein aggregates, and thus helps us analyze how the aggregation process occurs and how aggregates may play a role in these diseases. The paper is published open-access, so you should be able to read it to see all the details. (Naturally, the format of the mini-review limited the word count and meant that much had to be omitted)
Citation:
Patrick C.A. van der Wel; Solid-state nuclear magnetic resonance in the structural study of polyglutamine aggregation. Biochem Soc Trans 2024; BST20230731. doi: https://doi.org/10.1042/BST20230731
Publication: collaborative paper on perovskite-related materials, with Sn and Se ssNMR
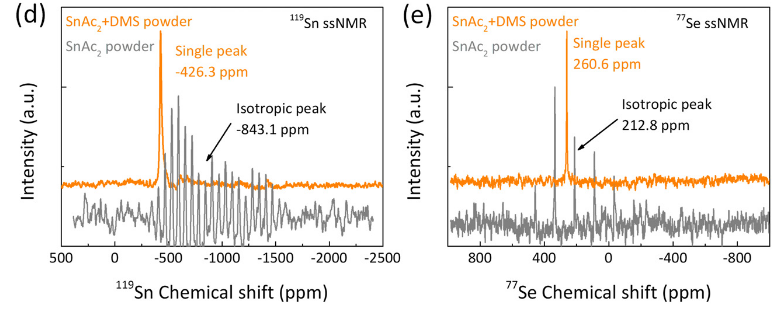
Congrats to our collaborators from the group of Maria Loi and others for the publication of their new paper on in situ SnSe deposition as passivation for scalable and stable quasi-2D lead–tin perovskite solar cells, in the journal Energy & Environmental Science. Dr. Lasorsa in our group contributed tin and selenium ssNMR analysis to this interdisciplinary paper. For more information see the paper at the journal.
Publication:
Chen L, Tekelenburg EK, Gahlot K, Pitaro M, Xi J, Lasorsa A, et al. In situ SnSe deposition as passivation for scalable and stable quasi-2D lead–tin perovskite solar cells. Energy Environ Sci. 2023;10.1039.D3EE02507A.
Publication: 13C labeling of hyaluronic acid polysaccharides (for ssNMR analysis)
Congrats to Pushpa and her collaborators on the new paper in the journal Carbohydrate Polymers. This new report describes Pushpa’s work to achieve the production of high-molecular-weight (HMW) hyaluronic acid, as part of her PhyCan (physics of cancer) project. Our interest in this polysaccharide stems from its important role in the extracellular matrix (ECM) of tissues, and in particular certain types of cancer tissues. In such tumors the amount of HA is upregulated, seemingly contributing to the progression of cancer development. A challenge in understanding this process is that HMW is difficult to study with most structural techniques, as it can be very large (megadaltons!) and highly dynamic. In this project, Pushpa outfitted HMW HA with 13C (and 15N) isotope labels, and shows that this makes multidimensional NMR (including solid-state NMR) feasible. This approach is expected to be valuable for many research areas, as HA is also important for many biomedical engineering (BME) type applications. ECM-mimicking hydrogels are for instance of great interest for engineering 3D environments for growing cells.
Publication:
Rampratap P, Lasorsa A, Perrone B, Van Der Wel PCA, Walvoort MTC. Production of isotopically enriched high molecular weight hyaluronic acid and characterization by solid-state NMR. Carbohydrate Polymers. 2023 Sep;316:121063.
Publication: Dynamics-based spectral editing to see (fiber) surfaces by solid-state NMR.
Congratulations to Dr. Irina Matlahov and (alum) Dr. Jennifer Boatz on the publication of their new paper in the Journal of Structural Biology X. The paper is entitled “Selective observation of semi-rigid non-core residues in dynamically complex mutant huntingtin protein fibrils“. It describes our latest research on the misfolded protein deposits associated with Huntington’s disease (HD), specifically looking at what happens on the surface of these protein fibrils.
In previous work we have studied the structure of these nanometer-sized fibrils formed by mutant huntingtin’s exon 1 fragment, using ssNMR, EM and other methods. In our earlier studies we used dynamics-sensitive NMR techniques to look at the core of these fibrils, taking advantage of its very rigid structure. Conversely, we had looked at super flexible protein parts that are away from the rigid core. Those types of techniques have been a popular tool for seeing rigid or flexible parts of (aggregated) proteins. However, in the current paper we try to look for protein parts with intermediate mobility. Of particular interest are those residues that form the surface of the rigid fibril core: residues that are somewhat immobilized by proximity to the core proper, but are somewhat mobilized by their interactions with water solvent surrounding the fibril. This part of the protein fibrils can be especially interesting, as it is the part seen by protein-targeting antibodies, PET ligands and protein-protein interaction partners in affected neurons.
The paper discusses how we can probe the impact of the water interactions with the surface through variable temperature ssNMR. However, it also shows the limitations of this traditional approach. Instead we advocate for a new type of dynamic filtering experiment that selectively shows the signals of semi-rigid/semi-mobile residues (such as those on the fiber surface). This technique (IMS-DYSE) is complementary to traditional DYSE methods that select rigid or flexible sites. For more details, please see the paper (linked below). We expect to combine this dynamic filtering technique with other types of pulse sequences, to probe in more detail the features of the fiber surface.
Note that this technique proved especially useful for our work on polyglutamine protein fibrils, as in this protein system we have a core made up of glutamine residues, providing an overwhelming strong NMR signal that masks the resonances of the surface glutamines. So, to see the surface, it is essential to find a way to suppress the core signals, leaving only those of the surface residues. The paper shows that this indeed works, and we can for the first time use this technique to distinguish the core and surface glutamines. With this technique in hand, we can foresee further studies of surface-specific features, such as the binding sites for targeted antibodies and/or amyloid-binding dyes.
The paper is available online at the journal, and has the following citation:
I. Matlahov, J.C. Boatz, P.C.A. van der Wel (2022) Selective observation of semi-rigid non-core residues in dynamically complex mutant huntingtin protein fibrils. J. Struct. Biol. X, vol. 6, 100077. DOI: 10.1016/j.yjsbx.2022.100077
PS. Some of the techniques and results from this publication were also discussed during a prior online seminar in the MIT ssNMR/DNP zoominar series. You can view that video here.
Publication: Collaborative paper with the Pescarmona group (ENTEG) on zeolites.
Congratulations to PhD student Mustapha El Hariri El Nokab and our collaborators from the Pescarmona group at ENTEG, on a new collaborative paper being accepted and posted online. In this work, Mustapha used both 29Si and 27Al magic angle spinning ssNMR to compare different zeolite samples, characterizing their chemical structure and degree of order. Aside from our ssNMR data, the paper features numerous other spectroscopic and synthetic methods. For more information, see the paper!
Zahra Asgar Pour, Romar Koelewijn, Mustapha El Hariri El Nokab, Patrick C. A. van der Wel, Khaled O. Sebakhy, Paolo Pescarmona (2022) Binder-free zeolite Beta beads with hierarchical porosity: synthesis and application as heterogeneous catalysts for anisole acylation. ChemCatChem in press.
Publication: SSNMR of alginate hydrogel (re)hydration
A new open-access publication by Mustapha and collaborators has been published in the journal Food Hydrocolloids, describing how he used various ssNMR measurements to probe alginate hydrogel structure and (re)hydration. Alginates can be cross-linked with calcium to form hydrogels, which are used in many different types of applications. This includes their use in slow-release drug delivery as well as food/nutritional applications. In this new paper, Mustapha shows how 1H, 2H and 13C ssNMR measurements can be used to see how these hydrogels are structured, but especially also how they are hydrated by the aqueous solvent. A key feature of the work is the use of 1H MAS NMR to detect the water molecules in the hydrogel macropores as well as those waters directly interacting with the alginate. The two types of water molecules form distinct ‘pools’ in the hydrogel, which can be nicely distinguished by their different molecular motion. The latter is detected via different relaxation measurements, in these NMR experiments.
To read more:
El Hariri El Nokab, M..; Lasorsa, A.; Sebakhy, K. O.; Picchioni, F.; van der Wel, P. C. A. Solid-State NMR Spectroscopy Insights for Resolving Different Water Pools in Alginate Hydrogels. Food Hydrocolloids 2022, 107500.
Publication: Structural and motional changes in a cytochrome c – lipid complex implicated in apoptosis.
Congratulations to lab alum Dr. Mingyue Li and our collaborators on the publication of a new paper in the Journal of Molecular Biology. The paper is online via its DOI link. The paper describes how we used solid-state NMR spectroscopy to characterise the partial destabilization of the native fold of the protein cytochrome c, as it is bound to cardiolipin lipids. This protein-lipid complex is implicated in the process of programmed cell death, where it plays a key role in triggering the self-destruction of undesired or disease cells in higher organisms. Notably, the CL-bound protein catalyses the process of mitochondrial lipid peroxidation, which we see being reconstituted in vitro via mass-spectrometry lipidomics. The latter work is done by our longstanding collaborators in the group of Valerian Kagan at the University of Pittsburgh. The current paper builds on our earlier work, but is of especial interest based on the fact that we bridge some of our prior findings (that suggested the protein to be surprisingly “folded” on the membrane) to studies that report a greater degree of mobility of the CL-bound protein. Here we see how the experimental conditions regulate protein mobility and also pinpoint how different extents of mobility are present in the membrane-bound cytochrome c. Interestingly, the regional dynamics map partly, but not completely onto the previously identified “foldons” that define the folding landscape of this widely studied mitochondrial protein. For more details see the paper below. It is available as #openaccess so just follow the DOI link for access:
Reference:
Mingyue Li, Wanyang Sun, Vladimir A. Tyurin, Maria DeLucia, Jinwoo Ahn, Valerian E. Kagan, Patrick C.A. van der Wel (2021) Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids, Journal of Molecular Biology, Volume 433, Issue 15, 23 July 2021, 167057
Publication: New paper on mitochondrial protein-lipid interactions published in PNAS.
Congratulations to lab alumns Dr. Abshishek Mandal and Dr. Jennifer Boatz, as well as our collaborators in the USA and Spain! A new collaborative paper on mitochondrial protein-lipid interactions has just been published in the journal PNAS. In this multidisciplinary work we studied how the protein Drp1 binds the special mitochondrial lipid cardiolipin in order to do its job managing the proper fission of mitochondrial membrane. Our collaborator Rajesh Ramachandran (at Case Western) coordinated a wide array of experimental and computational approaches to determine how Drp1’s “variable domain” (VD) binds cardiolipin. Along the way, two apparent CL-binding motifs were detected, which are seemingly shared by other CL-binding proteins. Their mutation disrupts CL binding and also the proper management of mitochondrial morphology, pointing to the importance of CL-based signals and interactions in this vital cellular process.
Experimentally, our group contributed various solid-state NMR measurements of the lipids, with which we probe the specificity of the interactions and also observe how the protein modulates the lipid bilayer itself. Moreover, we were involved in the sequence/structure analysis: sequence analysis of the “disordered” VD domain pinpointed those parts of the structure most likely to engage in lipid-driven folding upon membrane binding. The “MoRF” motifs (“molecular recognition features”) indeed appear to be involved in CL binding, based on NMR and mutational studies. In Drp1 they are intimately involved in a folding transition of the VD domain upon membrane binding. It will be interesting to see how widespread these CL-binding motifs (CBMs) are in other CL-binding proteins.
Reference:
[1] Mahajan M, Bharambe N, Shang Y, Lu B, Mandal A, Madan Mohan P, et al. NMR identification of a conserved Drp1 cardiolipin-binding motif essential for stress-induced mitochondrial fission. Proc Natl Acad Sci USA2021;118:e2023079118. https://doi.org/10.1073/pnas.2023079118.
Preprint: cytochrome c – cardiolipin studies by ssNMR on bioRxiv
Our latest update on structural studies of this peroxidase-active protein-lipid complex implicated in mitochondrial apoptosis has now posted to bioRxiv: https://www.biorxiv.org/content/10.1101/2021.02.24.432556v1
In the preprint we discuss how ssNMR reveals the involvement of specific and localised dynamics in the lipid-bound protein. Interestingly, the mobility is dependent on the bound lipid species, with an apparent correlation to the resulting peroxidase activity. The lipids thus act as both substrates and regulators of the pro-apoptotic enzymatic activity of the protein. This was also discussed in the recent webinar as discussed in an earlier post.
This work was made possible by several great collaborators at the University of Pittsburgh, and funding from the NIH for both the project and employed instrumentation.
Update: The paper has now been accepted for publication in the Journal of Molecular Biology, and can be found online at its DOI link.
Reference:
Mingyue Li, Wanyang Sun, Vladimir A. Tyurin, Maria DeLucia, Jinwoo Ahn, Valerian E. Kagan, Patrick C.A. van der Wel (2021) Activation of Cytochrome C Peroxidase Function Through Coordinated Foldon Loop Dynamics upon Interaction with Anionic Lipids, Journal of Molecular Biology, in press